library(haven)
library(ggplot2)
library(dplyr)
library(tidyr)
library(stringr)
library(mice)
library(ggcorrplot)
library(gtsummary)2 Generalized Estimating Equation
2.1 Data description
Potthoff and Roy growth data:
Growth measurements for 27 children (11 girls and 16 boys)
Distance (in mm) from the centre of the pituitary gland to the pterygo-maxillary fissure recorded at ages 8, 10, 12 and 14 years.
Interested in the differences between males and females in pattern of growth over the 4 time points (from age 8 to 14 years).
potthoffroy |>
pivot_longer(cols = -c(id,sex)) |>
mutate(age = str_extract(name,'\\d+') |> as.numeric()) |>
ggplot(aes(x = age,y = value))+
geom_line(aes(group = id))+
facet_wrap(~sex)+
theme_bw()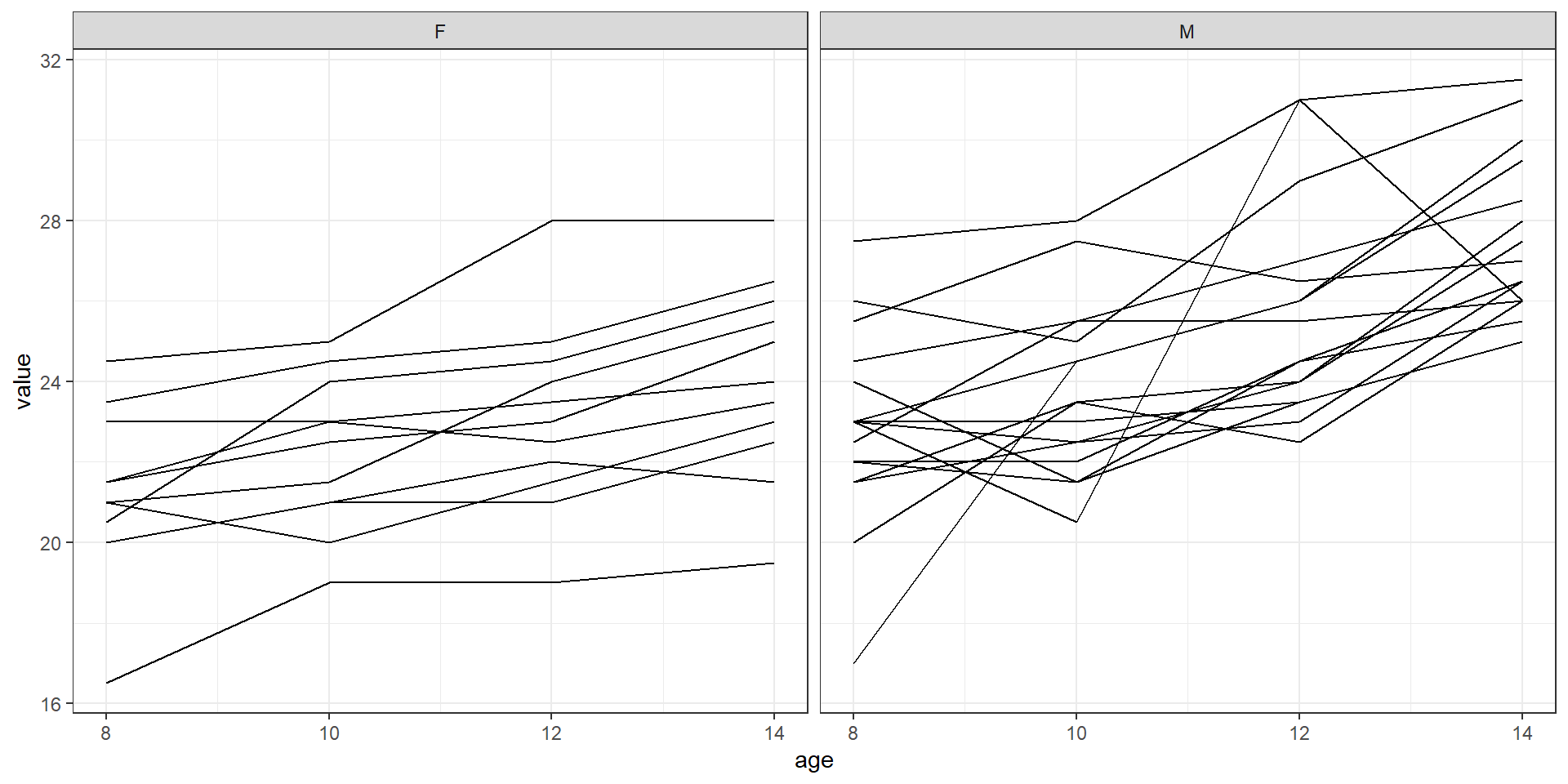
We consider the correlation of the value in male and female group
potthoffroy |>
filter(sex == "M") |>
select(-c(id,sex)) |>
cor(use="pairwise.complete.obs")|>
ggcorrplot(lab = TRUE) +
labs(tag = "Male")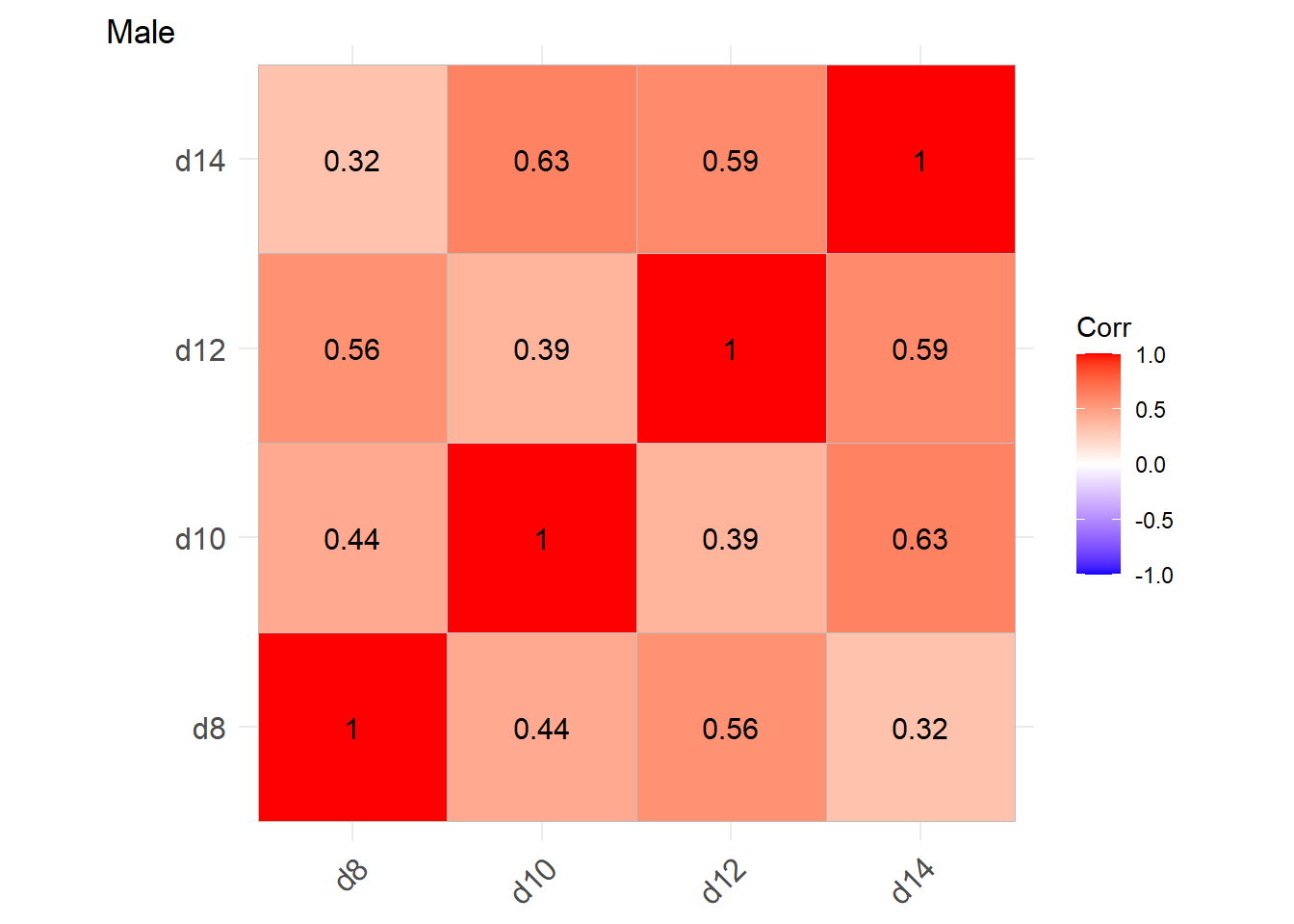
potthoffroy |>
filter(sex == "F") |>
select(-c(id,sex)) |>
cor(use="pairwise.complete.obs")|>
ggcorrplot(lab = TRUE) +
labs(tag = "Female")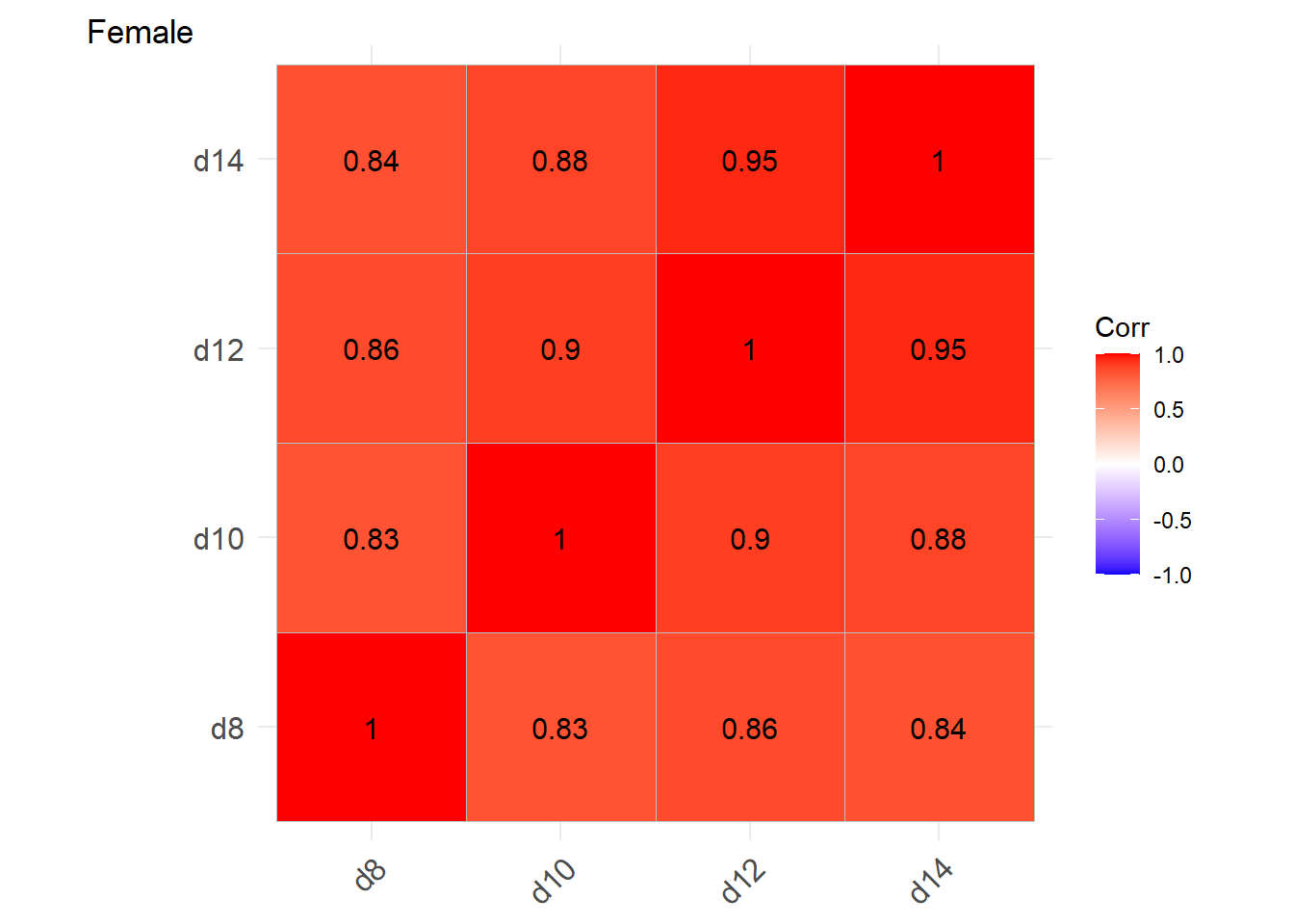
Positive correlations within-cluster. Greater for Females than Males.
2.2 Fitting OLS
data <- potthoffroy |>
pivot_longer(cols = -c(id,sex)) |>
mutate(age = str_extract(name,'\\d+') |> as.numeric(),
sex = factor(sex,labels = c(0,1)))
lm_model <- lm(value ~ sex + age, data = data)
summary(lm_model)
Call:
lm(formula = value ~ sex + age, data = data)
Residuals:
Min 1Q Median 3Q Max
-5.9882 -1.4882 -0.0586 1.1916 5.3711
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 15.38569 1.12857 13.633 < 2e-16 ***
sex1 2.32102 0.44489 5.217 9.20e-07 ***
age 0.66019 0.09776 6.753 8.25e-10 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 2.272 on 105 degrees of freedom
Multiple R-squared: 0.4095, Adjusted R-squared: 0.3983
F-statistic: 36.41 on 2 and 105 DF, p-value: 9.726e-13Pretend observations independent – WRONG!!
The estimation of beta (sex, age, intercept) is unbiased, but the standard errors are incorrect if the covariance is not independent.
\(\widehat{\beta}\) are estimated using Odinary Least Square
\(Var(\widehat{\beta})\) - derived from covariance matrix of \(Y\), known as V
Covariance = variance \(\times\) correlation
Conventional regression model fit by OLS has \(V_i = \sigma^2I_{n_i \times n_i}\)
2.3 How can we estimate the correct variance of the OLS estimator when the data are not independent?
Robust / empirical / “sandwich” variance estimator:
Use actual correlations observed in the data (i.e. empirical correlations)
Robust – for a large number of clusters it converges to the correct variance of the OLS parameter estimate, irrespective of what the true covariance V is, as long as the corrected model for the mean of Y has been specified.
Sandwich: the data driven empirical estimator is the “meat” in the sandwich between two model-based terms (“slices of bread”)
library(lmtest)
library(sandwich)
lm_model <- lm(value ~ sex + age, data = data)
## Robust step
cov_cl <- vcovCL(lm_model, cluster = data$id)
cov_cl (Intercept) sex1 age
(Intercept) 0.87447024 -0.3160789 -0.050590613
sex1 -0.31607886 0.5948969 -0.006492700
age -0.05059061 -0.0064927 0.005173734se_robust_model <- coeftest(lm_model, vcov = cov_cl)
se_robust_model
t test of coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 15.385690 0.935131 16.4530 < 2.2e-16 ***
sex1 2.321023 0.771296 3.0093 0.003279 **
age 0.660185 0.071929 9.1783 4.268e-15 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Compare
lm_model |>
tbl_regression(
intercept = TRUE,
estimate_fun = purrr::partial(style_ratio, digits = 3)) |>
remove_row_type(
type = "reference"
)| Characteristic | Beta | 95% CI | p-value |
|---|---|---|---|
| (Intercept) | 15.39 | 13.15, 17.62 | <0.001 |
| sex | |||
| 1 | 2.321 | 1.439, 3.203 | <0.001 |
| age | 0.660 | 0.466, 0.854 | <0.001 |
| Abbreviation: CI = Confidence Interval | |||
se_robust_model |>
tbl_regression(
intercept = TRUE,
estimate_fun = purrr::partial(style_ratio, digits = 3)
) | Characteristic | Beta | 95% CI | p-value |
|---|---|---|---|
| (Intercept) | 15.39 | 13.53, 17.24 | <0.001 |
| sex1 | 2.321 | 0.792, 3.850 | 0.003 |
| age | 0.660 | 0.518, 0.803 | <0.001 |
| Abbreviation: CI = Confidence Interval | |||
Sex: difference in mean outcome between sexes at any age = between-subject comparison
SE with robust > SE assuming independence (0.7712956 > 0.4448862)
Between cluster comparison is LESS precise with positive within-subject correlation
Age: effect estimated within-subject.
SE with robust < SE independence ( 0.0719287 < 0.0977589)
Within-cluster comparison is MORE precise with positive correlation
As we can see, within-cluster comparison is MORE precise with positive correlation. By modelling the covariance V correctly using the robust variance estimator instead of assuming independence, we generally obtain smaller standard errors. How to model covariance V?
2.4 Common correlation / covariance structures
Exchangeable: correlation between any two measurements on an individual is the same no matter how far apart in time they are.
Autoregressive: correlation decreases with distance
Unstructured: no pre-determined structure
Putting together in GEE
The covariance V is unknown, but take a “guest” at its form.
Call this W (e.g. exchangeable, autoregressive, etc.)
Can get idea from empirical correlations
Called a “working” covariance – we know it may be wrong but we work with it anyway
Step for GEE:
Fit OLS regression, and estimate parameters in W using residuals from OLS regression
Solve weighted least squares regression using \(\widehat{W}\) to get \(\widehat{\beta}_W\)
Iterate between \(\widehat{W}\) and \(\widehat{\beta}_W\) until convergence
2.5 Example data
Data from a study investigating recovery following appendectomy in children. The aim of the study was to determine whether the children who underwent a laparoscopic appendectomy achieved a faster rate of recovery than children who underwent conventional “open” appendectomy.
In this study a measure of recovery was made using an “uptimer”, a device worn by the children on their thigh. The uptimer senses the position of the thigh and records the times of changes from a horizontal position to a “vertical” position (at least 45 degrees from the horizontal). An increasing number of position changes (horizontal to vertical) from one day to the next is a marker of recovery.
appendix <- read_dta("./data/appendix.dta")
appendix <- appendix |>
mutate(lognchanges = log(nchanges),
group = factor(group,labels = c("Lap","Open"))) Plot the distribution of number of change
appendix |>
ggplot(aes(nchanges))+
geom_histogram()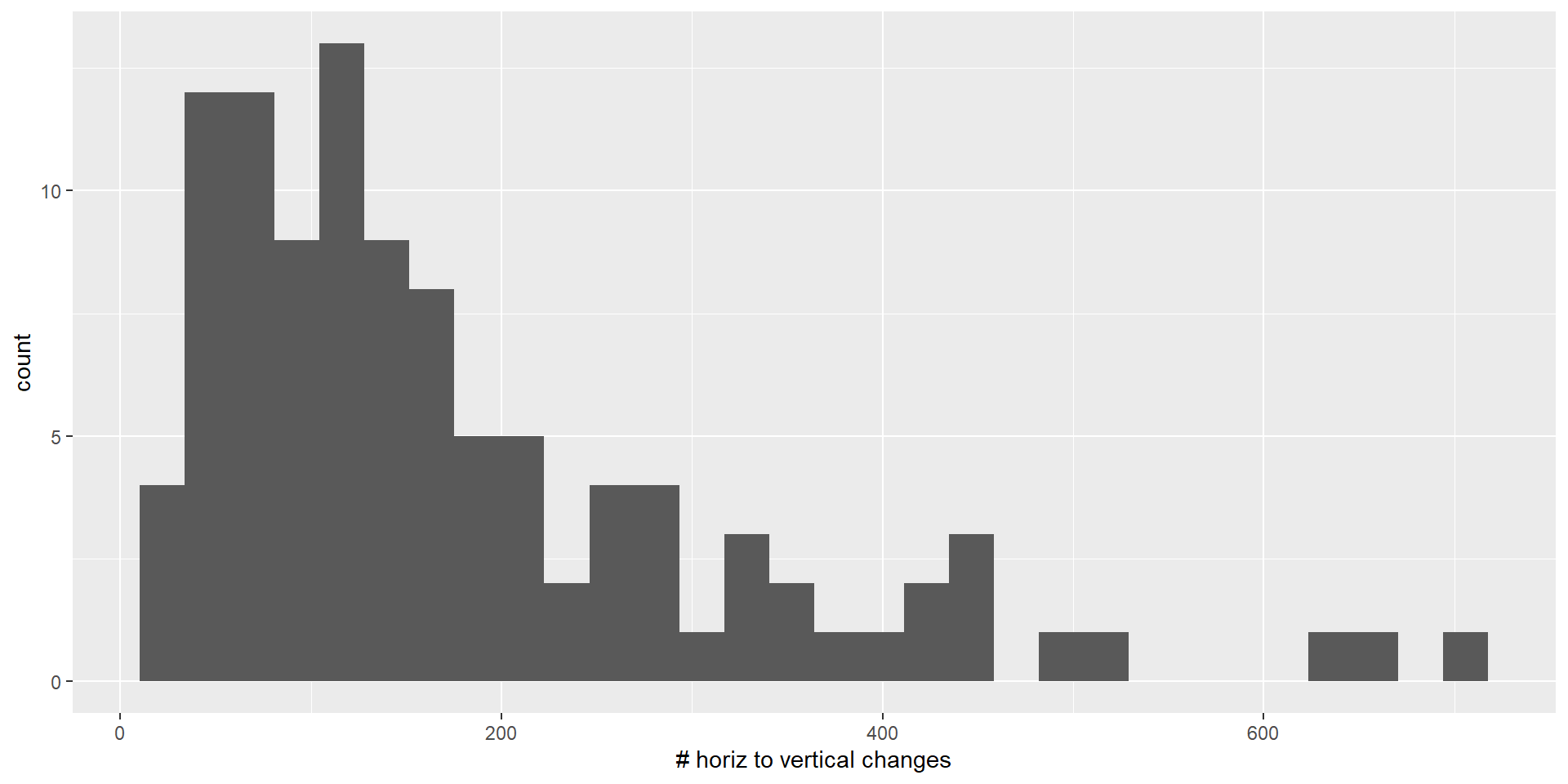
This seem like a skewed distribution, so we will use logarithm
appendix |>
ggplot(aes(lognchanges))+
geom_histogram()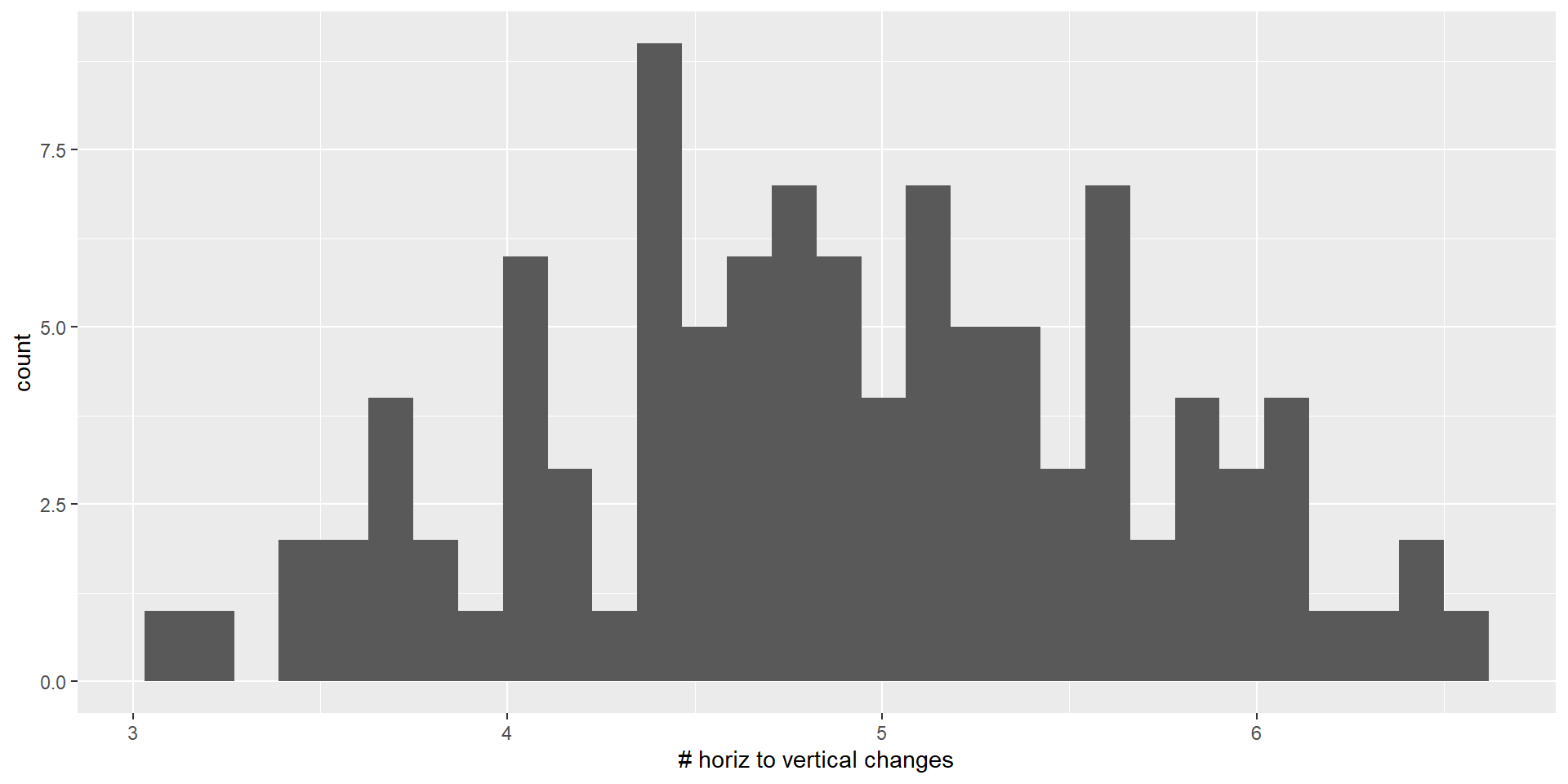
And we will see the description of the data
appendix |>
ggplot(aes(x = day, y = lognchanges))+
geom_line(aes(group = patid))+
geom_point()+
facet_wrap(~group)+
theme_bw()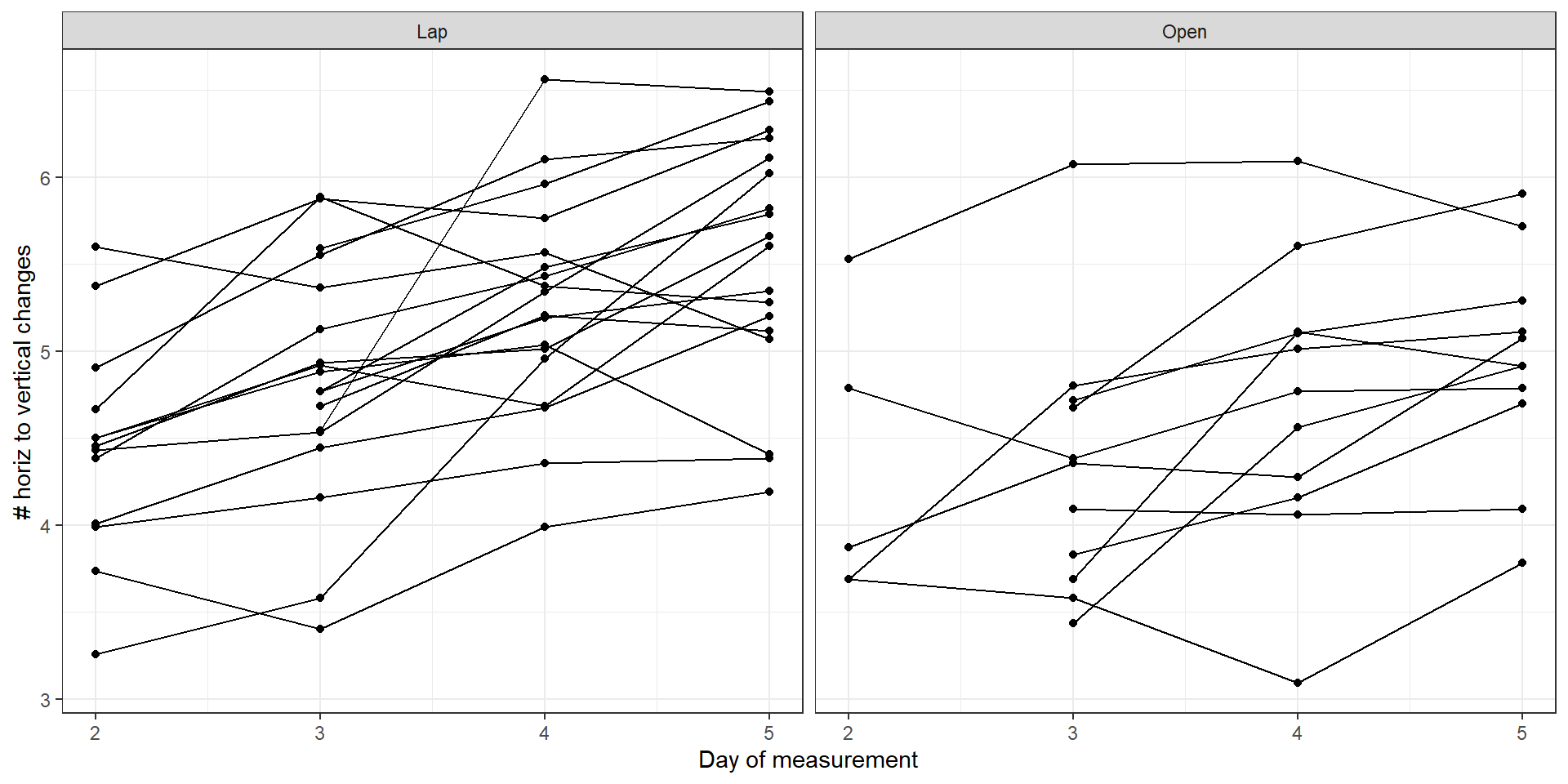
Using GEE with different working covariance \(W\)
The research question is whether the rate of change of the outcome variable – logged number of position changes – differs between the two groups. In order to address this within a regression modeling framework, we need to model the relationship between the outcome and time, and allow this relationship to differ between groups. The effect of surgical group will then be measured by a single interaction parameter.
library(geepack); library(gee)geefit.ind <- geeglm(lognchanges~day*group,
data = appendix,
id = patid,
corstr = "independence")
summary(geefit.ind)
Call:
geeglm(formula = lognchanges ~ day * group, data = appendix,
id = patid, corstr = "independence")
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) 3.74588 0.29367 162.700 < 2e-16 ***
day 0.36338 0.07159 25.763 3.86e-07 ***
groupOpen -0.04868 0.50811 0.009 0.924
day:groupOpen -0.11807 0.11090 1.133 0.287
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = independence
Estimated Scale Parameters:
Estimate Std.err
(Intercept) 0.4488 0.08769
Number of clusters: 29 Maximum cluster size: 4 geefit.ex <- geeglm(lognchanges~day*group,
data = appendix,
id = patid,
corstr = "exchangeable")
summary(geefit.ex)
Call:
geeglm(formula = lognchanges ~ day * group, data = appendix,
id = patid, corstr = "exchangeable")
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) 3.8441 0.2724 199.13 < 2e-16 ***
day 0.3407 0.0676 25.41 4.6e-07 ***
groupOpen -0.2163 0.4333 0.25 0.62
day:groupOpen -0.0794 0.1007 0.62 0.43
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = exchangeable
Estimated Scale Parameters:
Estimate Std.err
(Intercept) 0.449 0.088
Link = identity
Estimated Correlation Parameters:
Estimate Std.err
alpha 0.668 0.0988
Number of clusters: 29 Maximum cluster size: 4 geefit.ar1 <- geeglm(lognchanges~day*group,
data = appendix,
id = patid,
corstr = "ar1",wave=day)
summary(geefit.ar1)
Call:
geeglm(formula = lognchanges ~ day * group, data = appendix,
id = patid, waves = day, corstr = "ar1")
Coefficients:
Estimate Std.err Wald Pr(>|W|)
(Intercept) 3.7648 0.2494 227.89 < 2e-16 ***
day 0.3539 0.0656 29.12 6.8e-08 ***
groupOpen -0.2869 0.4374 0.43 0.51
day:groupOpen -0.0613 0.1015 0.36 0.55
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation structure = ar1
Estimated Scale Parameters:
Estimate Std.err
(Intercept) 0.45 0.0882
Link = identity
Estimated Correlation Parameters:
Estimate Std.err
alpha 0.76 0.079
Number of clusters: 29 Maximum cluster size: 4 | Independence | Exchangeable | Auto-regressive 1 | |
|---|---|---|---|
| Group effect (on slope) | -0.1181 | -0.0794 | -0.0613 |
| Robust SE of group effect (on slope) | 0.1109 | 0.1007 | 0.1015 |